Learn Python
This page is an invitation to learn Python and apply it to bioinformatics. It is designed for people with no programming experience who are interested in the possibility of learning how to program. It uses my limited experience in this area to demonstrate that even if you were never exposed to any programming it is possible to learn enough Python to write programs that analyze sequence data and produce results. In addition, programming is fun and it beats Sudoku and crossword puzzles as a constructive brain teaser and pastime. -Luca Comai
What is Python?
Python is a modern programming language that is easy to code and use. It resembles another very common language called Perl. Python code is easy to understand because the language resembles natural language and because it uses indentation to separate blocks of code and to convey their hierarchy. Python is very powerful and is used by major private and public institutions such as Google and NASA.
How I learned
I first tried Perl, another programming language, and became frustrated by my inability to understand the syntax. Now that I know a little bit about programming, I am not clear why this is the case because I can understand (kind of) Perl and it is really not difficult. This just goes to show that learning programming may seem difficult at first. When that happened, I dropped the effort. A few months later, I decided to try again. Victor Missirian, a bioinformatician who worked with me, showed me the Python tutorial written by the Python inventor, Guido von Rossum. It appeared simple to follow and so I decided to try again. I downloaded the Python installation package from python.org, bought a couple of books and never looked back. I started writing a program that would count and report all the restriction fragments in a genome. Having a specific objective helped me focusing and motivating me. Since then I have written dozens of little programs to do all kinds of stuff, such as parsing Illumina sequencing files, grading my class, performing in silico comparative genomic hybridization, analyzing the genomic outcome of crosses, and so on. Now, do not get me wrong, I am not sure I would call myself an expert and there is a lot left for me to learn. But this is really the good news. You do not need a degree in computer sciences to have fun and be productive.
How you can learn
Get Python
Apple computers come with a version of Python installed. It is useful, however, to download a Python as part of a software package manager called Anaconda. You could get a Python installer from the official Python website, but Anaconda is handy because it installs a set of auxiliary programs for data science. Python comes in two versions: the old one is 2.x, the new one is 3.x. I suggest you download the latest 3.x, such as 3.8 (June 2020). Anaconda provides an Interactive Developer Environment (IDE) called Spyder, in which you can write a program and execute it. There are many other IDE available. Another way to write and execute a program is a code editor. It is similar to an IDE, but more sophisticated in handling text. A very good one is BBedit (free version available). Last, you can run Python inside a notebook, a very useful method that will be discussed in a different post.
Start practicing
With Python open in an IDE or in the Terminal (for Apple computers), follow the Python tutorial . Once I learned the basics, I found two books very useful. The first is Learning Python by Mark Lutz. The second is Python Cookbook by B.K. Jones. Beyond the books, the Internet has an answer for just about any problem you may encounter: just google it. By the way, while you are working on your skills it would be good to also learn basic Unix commands: see this tutorial.
Examples of programs
To download a program go the program page using the link below, then click the download link. Depending on what you did the two following outcomes are possible.
- If your browser takes you to a new page, select all the text in the page and copy it into IDE or Text Wrangler. Save it by adding a “.py” extension. The program will color the code to distinguish annotation from code.
- If you right-button click on the download link, you should be able to download the program as a “.py” file.
Note that the program contains both “live” code and annotation. The latter is any line that is flanked by triple apostrophes: “‘ “‘ , and any line that starts with “#”. The annotation is there to explain the program or to remind the author or user about the function for each line of code.
Barcode generator
Barcode generator produces DNA barcodes of desired length and sequence distance for use in adapters for sequencing libraries .
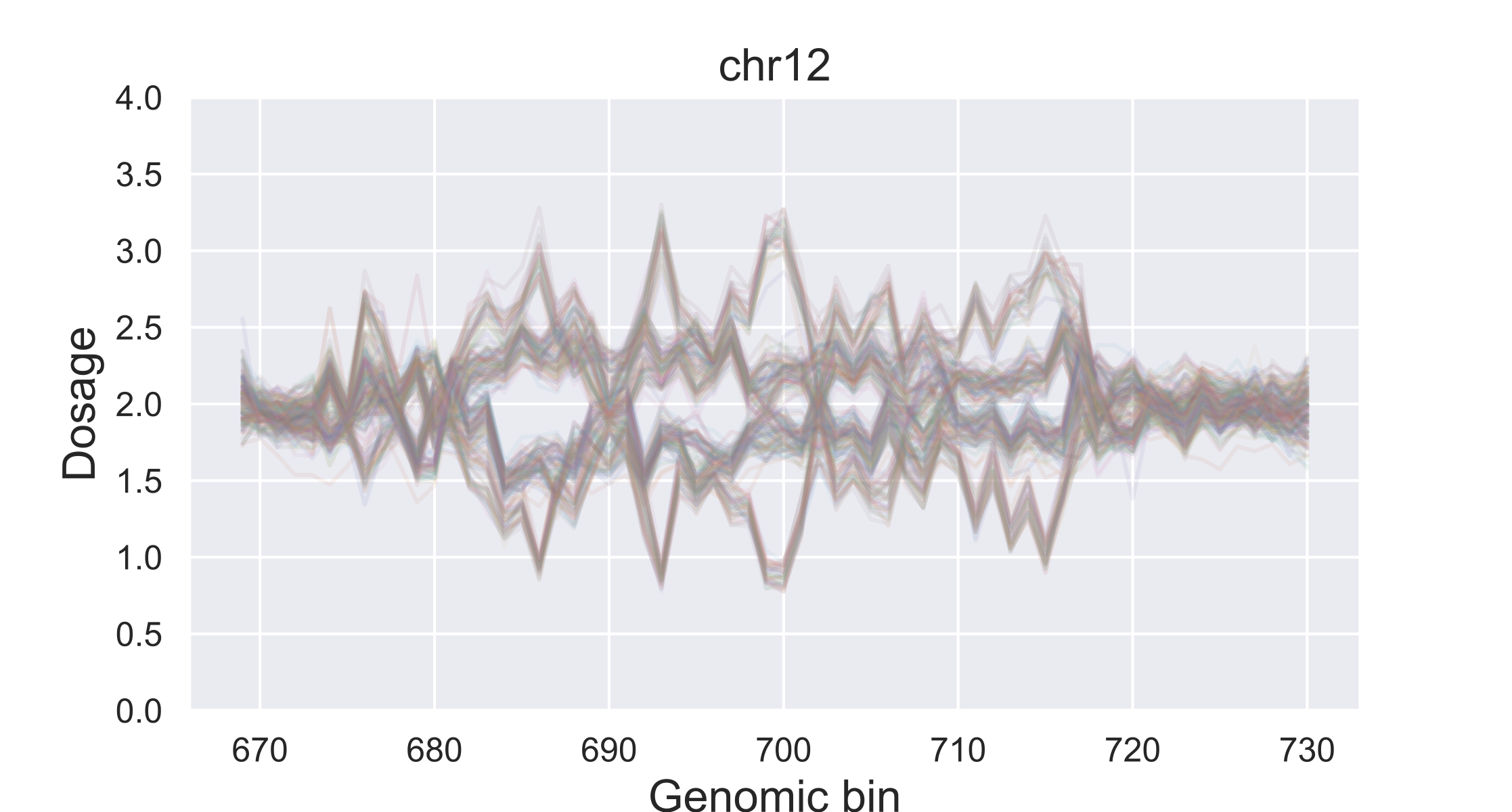
CNV Mapping
CNV Mapping uses mapped sequencing reads from related individuals in a population to derive linkage between copy number variable loci. This program is written as a notebook: it is divided in cells within a program called Jupyter Notebook, also part of the Anaconda installation. The notebook format has great utility, which I will explain in another post.
Funding sources
The research experience on which the Python pages are based is funded by the National Science Foundation Plant Genome grant DBI-0733857 (Functional Genomics of Polyploids), NSF Plant Genome award DBI-0822383, TRPGR: Efficient identification of induced mutations in crop species by ultra-high-throughput DNA sequencing, and National Institutes of Health R01 GM076103-01A1 (Dosage dependent regulation in hybridization) to LC.

this is the very easy way of learning for python coures i have see this blog reseanty
LikeLike